This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein domains?
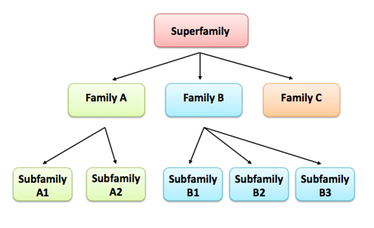
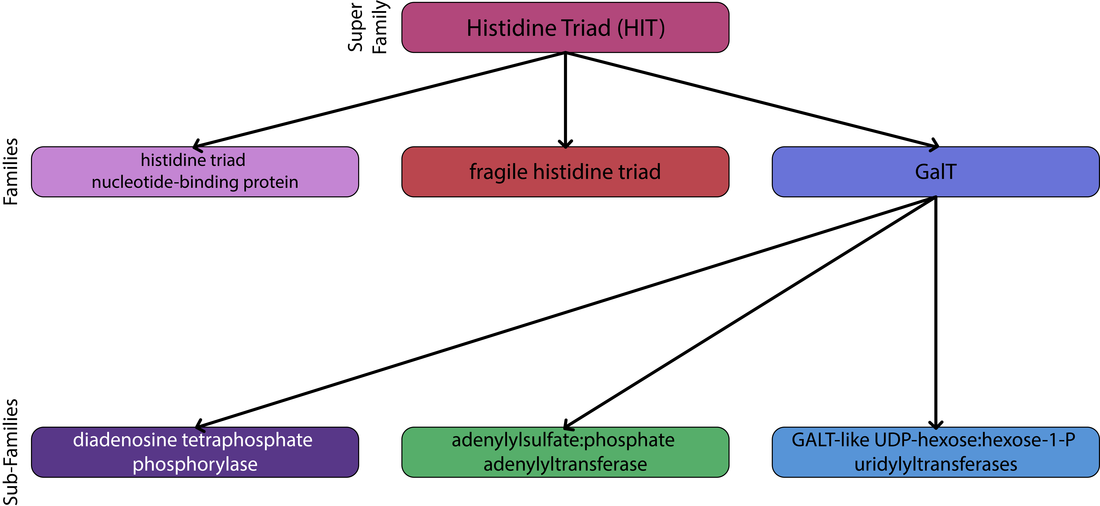
Domains are distinct functional and structural units responsible for a particular function or interaction, which define the proteins overall role [1]. Domains are also evolutionary conserved portions of a protein sequence that can evolve, function, and exist independently of the rest of the protein. Thus proteins can be classified based on sequence or structural similarity, which are commonly referred to as protein families. Families are defined as proteins which share a common evolutionary history, which is reflected by their shared function or similarity in sequence [1]. Proteins are typically arranged in a hierarchical structure (superfamilies, families, and subfamilies). A superfamily being a large group of distantly related proteins and a subfamily a small group a closely related proteins (Fig. 1)
What domains are found in GALT?
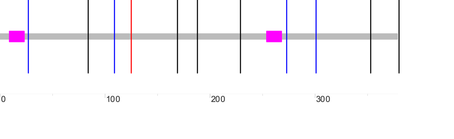
I used the online programs Pfam, SMART, InterPRo, and PROSITE to search the Human GALT protein for domains. This was done to compare outputs to ensure I get consistent and similar results. The results are displayed below:
Both Pfam and InterPro identified two domains within GALT, while PROSITE identified one domain, and SMART identified no domains. It was expected that SMART would not identify any domains because its algorithm was deliberately chosen to ignore domains involved in metabolism (since these domains are typically longer and very well conserved ). PROSITE only identified the active site of GALT.
So what are the two domains?
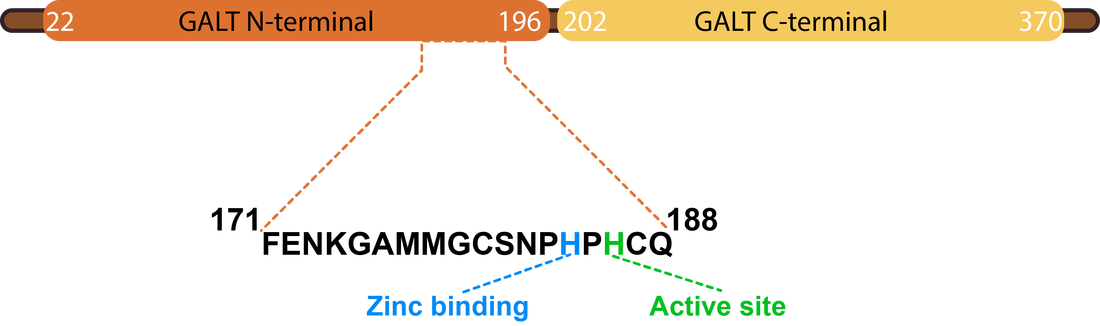
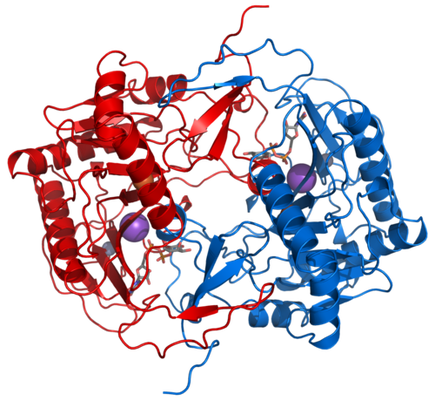
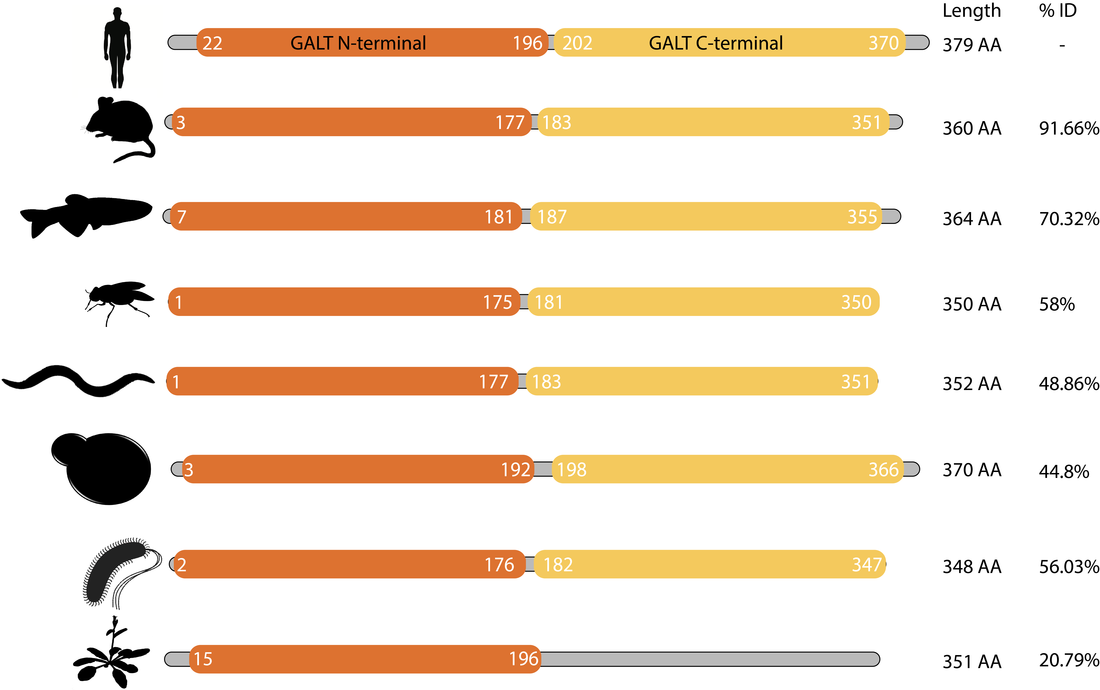
Both Pfam and InterPro identified two domains, GALT N-terminal and GALT C-terminal domains. The GALT N-terminal domain contains the active site of the GALT enzyme, which catalyzes the conversion of UDP-glucose and alpha-D-galactose 1-phosphate to alpha-D-glucose 1-phosphate and UDP-galactose [2]. The role of the structure of this domain seems to be to present the hexose-phosphate to the active site (histidine) while excluding competing reactants (water). The GALT C-terminal domain has been reported to have been formed by a fold-duplication of the N-terminal domain. Both domains are involved in the binding of a zinc and iron atom [3]. The active version of the GALT enzyme is a dimer, formed by two identical GALT proteins (see below).
How conserved are the domains of GALT?
The domain structure of the GALT protein is well conserved from mammals to bacteria. Interestingly, the GALT of Arabidopsis thaliana did not have a detectible GALT C-terminal domain.
How conserved is the active site of GALT?
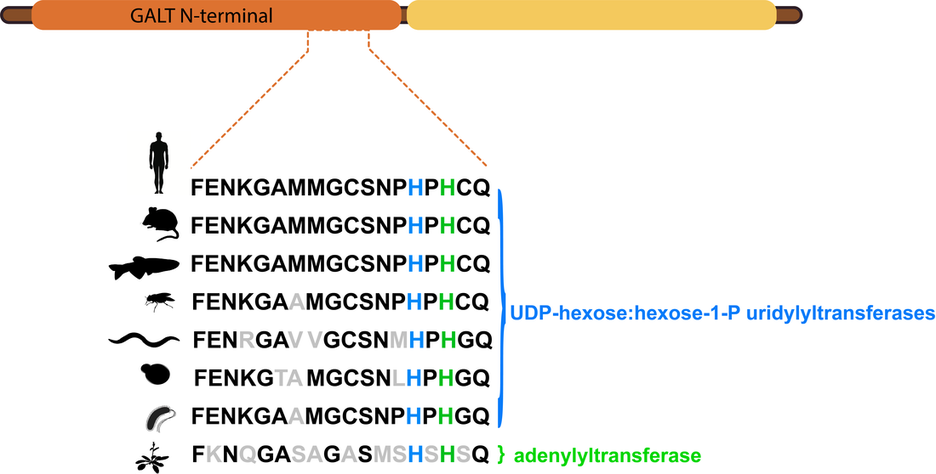
The active site of GALT is absolutely critical for it to function properly. In particular, the motiff HxHxQ is essential for the formation of active site and catalysis of UDP-glucose and alpha-D-galactose 1-phosphate to alpha-D-glucose 1-phosphate and UDP-galactose [3]. The first Histidine is involved in binding zinc, which contributes to the stability of the active site, while the second Histidine acts as the nucleophile in catalysis [3].
What superfamily are GALT proteins a part of?
Based on the above analysis it is clear that the GALT of Arabidopsis thaliana (NP_197321.1) is clearly not a close homolog of the other GALT proteins. In fact an earlier study solved the structure of this protein and identified that it is in fact an adenylyltransferase. But, due to its similar sequence to GALT, it was annotated as such [4]. While this protein is not a close homolog of GALT it is a distant relative, belonging to the same protein family. Below depicts the family structure of GALT and related proteins.
The Human GALT protein belongs to the histidine triad super-family of nucleotide hydrolases and transferases and the GALT-like UDP-hexose:hexose-1-P uridylyltransferases sub-family. This super family houses three families: the histidine triad nucleotide-binding protein family, the fragile histidine triad family, and the GalT family [5]. The GalT family is distinct from the other two in the following regards: it does not have a triad of histidines, but a conserved HxHxQ sequence, and the reactions carried out by proteins in this family are nucleotidyl transfer rather than hydrolysis [6]. It should be noted that the most common Human disease allele occurs at the glutamine in this highly conserved site.
Conclusions
From Humans to E. coli, GALT has to two domains which aid in catalysis and binding of metal atoms. Further, these proteins are highly conserved at the active site, with a conserved motif (HxHxQ) involved in catalysis and active site stability. Upon further review, the annotated GALT of Arabidopsis thaliana is actually a more distantly related protein with adenylyltranferase activity. But, nonetheless is a member of the conserved Galt family.
References
1. Protein classification: An introduction to EMBL-EBI resources
2. Wedekind et al. The Structure of Nucleotidylated Histidine-166 of Galactose-1-phosphate Uridylyltransferase Provides Insight into Phosphoryl Group Transfer. Biochemistry 1996;35:11560-11569
3. Wedekind et al. Three-Dimensional Structure of Galactose- 1 -phosphate Uridylyltransferase from Escherichia coli at 1.8 A Resolution. Biochemistry 1995;34:11049- 11061
4. McCoy et al. Structure and Mechanism of an ADP-Glucose Phosphorylase from Arabidopsis thaliana. Biochemistry 2006;45:3154–3162
5. Brenner et al. Crystal structures of HINT demonstrate that histidine triad proteins are GalT-related nucleotide-binding proteins. Nat. Struct. Biol. 1997;4:231–238.
6. Brenner et al. GalT: function, structure, evolution, and mechanism of three branches of the histidine triad superfamily of nucleotide hydrolases and transferases. Biochemistry. 2002;41:9003–9014.
2. Wedekind et al. The Structure of Nucleotidylated Histidine-166 of Galactose-1-phosphate Uridylyltransferase Provides Insight into Phosphoryl Group Transfer. Biochemistry 1996;35:11560-11569
3. Wedekind et al. Three-Dimensional Structure of Galactose- 1 -phosphate Uridylyltransferase from Escherichia coli at 1.8 A Resolution. Biochemistry 1995;34:11049- 11061
4. McCoy et al. Structure and Mechanism of an ADP-Glucose Phosphorylase from Arabidopsis thaliana. Biochemistry 2006;45:3154–3162
5. Brenner et al. Crystal structures of HINT demonstrate that histidine triad proteins are GalT-related nucleotide-binding proteins. Nat. Struct. Biol. 1997;4:231–238.
6. Brenner et al. GalT: function, structure, evolution, and mechanism of three branches of the histidine triad superfamily of nucleotide hydrolases and transferases. Biochemistry. 2002;41:9003–9014.
Figures
Figure 1. http://www.ebi.ac.uk/training/online/sites/ebi.ac.uk.training.online/files/figure2.png
Figure 3. https://en.wikipedia.org/wiki/File:Galactose-1-phosphate_uridylyltransferase_1GUP.png
Figure 3. https://en.wikipedia.org/wiki/File:Galactose-1-phosphate_uridylyltransferase_1GUP.png